2012年4月30日星期一
2012年4月27日星期五
Nexus data format and Newick tree format
1. Nexus data format for phylogenetic analysis
http://en.wikipedia.org/wiki/Nexus_file
2. Newick tree format for phylogenetic tree
http://en.wikipedia.org/wiki/Newick_format
http://en.wikipedia.org/wiki/Nexus_file
2. Newick tree format for phylogenetic tree
http://en.wikipedia.org/wiki/Newick_format
implications for improving the quality of seeds - 系统进化研究的现实意义
decipher evolutionary innovations - 解码进化创新
positive emotions and wellbeing - 好情绪与幸福
Positive emotions not only contribute to our success, they don't just contribute to our feeling good; they also contribute to our wellbeing.
积极情绪不仅有助成功,不仅能让我们感觉好,还有助我们获得幸福。
2012年4月26日星期四
2012年4月25日星期三
2012年4月24日星期二
2012年4月23日星期一
Plant genome diversity - a springer book
http://www.springerlink.com/content/978-3-7091-1129-1/contents/
2012年4月22日星期日
2012年4月20日星期五
what makes or breaks a Phd student
http://bioinformatics.psb.ugent.be/jobs
Nature published a short career article about what makes or breaks a Phd student . We think the profile below matches our Phd students quite well . Does it match you ?
We'd like to add one more thing to the list :
- Choose a supervisor whose work you admire and who is well supported by grants and departmental infrastructure.
- Take responsibility for your project.
- Work hard
: long days all week and part of most weekends .If research is your passion this should be easy ,and if it isn't ,you are probably in the wrong field .Note who goes home with a full briefcase to work on at the end of the day .This is a cause of success ,not a consequence . - Take some weekends off, and decent holidays, so you don't burn out.
- Read the literature in your immediate area, both current and past, and around it. You can't possibly make an original contribution to the literature unless you know what is already there.
- Plan your days and weeks carefully to dovetail experiments so that you have a minimum amount of downtime.
- Be creative : Think about what you are doing and why, and look for better ways to go. Don't see your PhD as just a road map laid out by your supervisor.
- Develop good writing skills : they will make your scientific career immeasurably easier.
To be successful you must be at least four of the following: smart, motivated, creative, hard-working, skillfuland lucky .You can't depend on luck ,so you should better focus on the others !
It's better to burn out than to fade away. Neil Young
2012年4月19日星期四
2012年4月18日星期三
data set from a re-sequencing project of rice
Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes
ftp://public.genomics.org.cn/BGI/rice/rice_resequencing/02.SNP/50_total/individual.txt
1 01 45975_AUS 45975_AUS C AUS 2 02 6307_AUS 6307_AUS C AUS 3 03 27762_IND 27762_IND C IND 4 04 30416_IND 30416_IND C IND 5 05 43545_IND 43545_IND C IND 6 06 51250_IND 51250_IND C IND 7 07 51300_IND 51300_IND C IND 8 08 8231_IND 8231_IND C IND 9 09 9148_IND 9148_IND C IND 10 10 9177_IND 9177_IND C IND 11 11 25901_IND 25901_IND C IND 12 12 6513_IND 6513_IND C IND 13 13 12883_AUS 12883_AUS C AUS 14 14 8555_AUS 8555_AUS C AUS 15 15 1107_TEJ 1107_TEJ C TEJ 16 16 27630_TEJ 27630_TEJ C TEJ 17 17 32399_TEJ 32399_TEJ C TEJ 18 18 2540_TEJ 2540_TEJ C TEJ 19 19 418_TEJ 418_TEJ C TEJ 20 20 55471_TEJ 55471_TEJ C TEJ 21 21 8191_TEJ 8191_TEJ C TEJ 22 22 NP_TEJ NP_TEJ C TEJ 23 23 11010_TRJ 11010_TRJ C TRJ 24 24 17757_TRJ 17757_TRJ C TRJ 25 25 328_TRJ 328_TRJ C TRJ 26 26 38698_TRJ 38698_TRJ C TRJ 27 27 43325_TRJ 43325_TRJ C TRJ 28 28 26872_TRJ 26872_TRJ C TRJ 29 29 43675_TRJ 43675_TRJ C TRJ 30 30 50448_TRJ 50448_TRJ C TRJ 31 31 66756_TRJ 66756_TRJ C TRJ 32 32 8244_TRJ 8244_TRJ C TRJ 33 33 12793_ARO 12793_ARO C ARO 34 34 38994_ARO 38994_ARO C ARO 35 35 9060_ARO 9060_ARO C ARO 36 36 9062_ARO 9062_ARO C ARO 37 37 RA4952_ARO RA4952_ARO C ARO 38 38 31856_ARO 31856_ARO C ARO 39 39 43397_TRJ 43397_TRJ C TRJ 40 40 60542_IV 60542_IV C IV 41 41 nivara_105327 N_105327 W N 42 42 nivara_106105 N_106105 W N 43 43 nivara_106154 N_106154 W N 44 44 nivara_80470 N_80470 W N 45 45 nivara_89215 N_89215 W N 46 46 rufipogon_105958 R_105958 W R 47 47 rufipogon_105960 R_105960 W R 48 48 rufipogon_Nepal R_Nepal W R 49 49 rufipogon_P46 R_P46 W R 50 50 rufipogon_YJ R_YJ W R
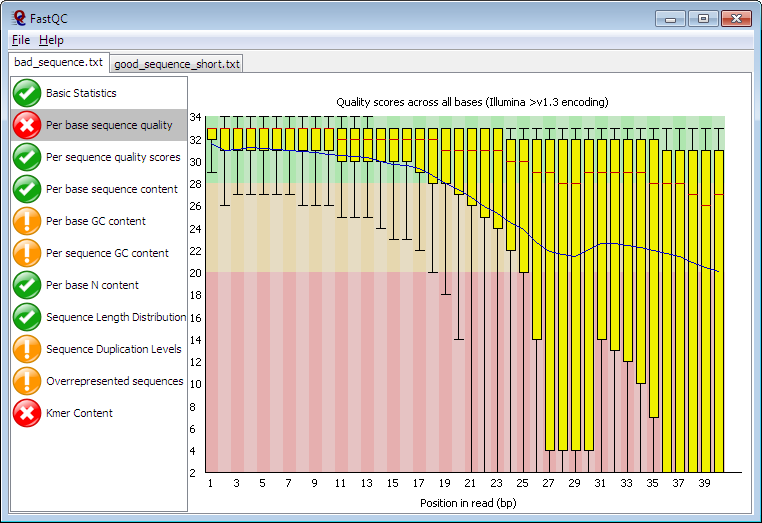
FastQC - a simple way to do some quality control checks on NGS data
http://www.bioinformatics.babraham.ac.uk/projects/fastqc/
FastQC aims to provide a simple way to do some quality control checks on raw sequence data coming from high throughput sequencing pipelines . It provides a modular set of analyses which you can use to give a quick impression of whether your data has any problems of which you should be aware before doing any further analysis .
The main functions of FastQC are
- Import of data from BAM, SAM or FastQ files (any variant)
- Providing a quick overview to tell you in which areas there may be problems
- Summary graphs and tables to quickly assess your data
- Export of results to an HTML based permanent report
Offline operation to allow automated generation of reports without running the interactive application
high level of creativity and depressed 创新与抑郁
If you want to have high level of creativity, it's a must you have to be depressed.
如果你想拥有高水平的创造力,就必须承受抑郁。
Analyzing Biological Data Using R: Methods for Graphs and Networks
http://www.springerprotocols.com/Abstract/doi/10.1007/978-1-61779-361-5_19
R is a powerful language and widely used software tool for the analysis and visualization of data . Its core capabilities can be extended through many different add-on packages . Among the many packages are some which offer a broad range of facilities for analyzing statistical properties of graphs . This chapter provides a practical tutorial covering the use of R methods for graphs and networks to examine biological data and analyze their topological and statistical properties .
2012年4月17日星期二
perform evolutionary or population-based analyses directly on BAM/SAM files
POPBAM, http://popbam.sourceforge.net/
POPBAM is a tool to perform evolutionary or population-based analyses of next-generation sequencing data . POPBAM takes a BAM file as its input and can compute many widely used evolutionary genetics measures in sliding windows across a genome . The motivation for developing POPBAM is to provide the community with a fundamental suite of evolutionary analyses tools that would otherwise be tedious to implement . Since POPBAM works directly with BAM files , there are no intermediary steps necessary .
To enable POPBAM to perform population-level analyses, it is first necessary to modify the input BAM file header. Users must add the "PO" tag to the header line for each read group. The "PO" tag can be any string, as long as the string is identical between samples from the same population. One example may be that a BAM file has three read groups (R21, R22, and R25). The R22 and R25 read groups are from two different lines of Drosophila melanogaster called "MEL01" and "MEL02", while the third read group, R21, is from a single line of D. simulans called "SIM01". Below is an example of the BAM header including the "PO" tag:
@RG ID:R22 SM:MEL01 PO:MEL @RG ID:R25 SM:MEL02 PO:MEL @RG ID:R21 SM:SIM01 PO:SIM
2012年4月15日星期日
2012年4月13日星期五
DnaSAM - DNA Sequence Analysis and Manipulation
DnaSAM - DNA Sequence Analysis and Manipulation
订阅:
博文 (Atom)